Automating Molecular Diagnostics
Viapath has begun working with diagnostics.ai to implement artificial intelligence into its laboratory’s molecular assays. The technology automates the analysis and reporting of results, with built-in quality control, ensuring accuracy and standardisation.
The innovative system removes the manual steps associated with analysing the data, delivering a digitised and paperless system (tying into the NHS Five Year Forward View - Paperless 2020), as well as enabling faster turnaround times, reduced risk and improved laboratory capacity.
The initial laboratory implementation will be in Viapath’s South London Specialist Virology Centre at King’s College Hospital. The first systems to be automated will be the high-volume real-time PCR testing, used for a wide range of sample types and tests including respiratory, STD and gastrointestinal assays through the use of diagnostics.ai’s cloud-based service, pcr.ai. The implementation began in February 2018, demonstrating Viapath’s continued commitment to setting the standard for the future of pathology.
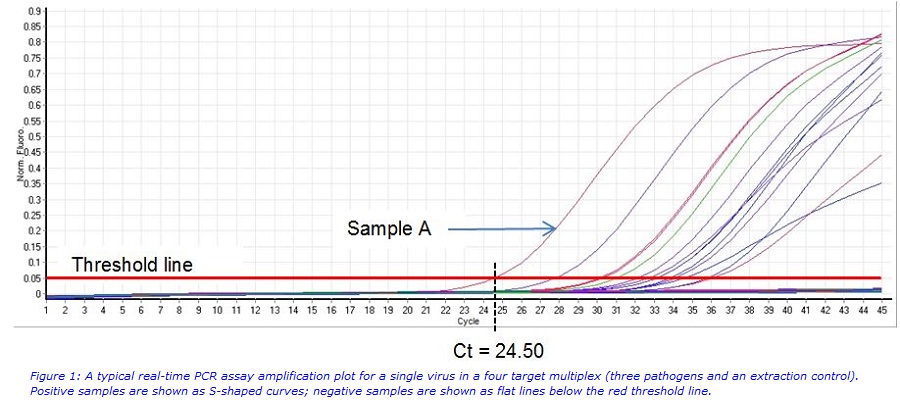
To illustrate the complexities of manually analysing real time PCR assay data, figure 1 shows a typical set of amplification curves. The instrument’s software plots the amplification of viral nucleic acid during the reaction based on the increase in signal from fluorescent probes, specific to the viral target. During the amplification reaction more and more copies of the target are made, allowing more probes to bind resulting in an increase in the fluorescent signal. A sample is positive if an S-shaped curve is produced that crosses the red threshold line. This results in a Ct value (the cycle number on the X-axis). In figure 1 sample A is positive with a Ct value of approximately 24.5.
The example above comes from a multiplex assay that detects three viruses together with an extraction control. This means that there are four separate amplification plots that need to be checked manually for each run. This can be a time consuming process, particularly if there are any difficult to interpret amplification traces. This will require further, time-consuming analyses which can involve re-setting reaction parameters such as baseline and threshold values, or noise-band ranges to determine which samples are positive and which are negative.
Furthermore, quality assurance data has to be entered for the standards for each target together with values for the extraction control, which is used to monitor each sample for reaction inhibition. This is to ensure that if a sample is negative, it is a true negative and not because the reaction has failed due to an inhibitor carried over during the extraction process. Setting reaction parameters and visual interpretation of results is by definition subjective and requires considerable time, expertise and quality control procedures. In contrast, a patented machine learning algorithm that automatically interprets both quantitative and qualitative real-time PCR tests and digitises the results without manipulating the raw data has been developed by diagnostics.ai. This means that there is no need for manual interpretation, or to look at the amplification traces to analyse the results, although the software allows the user to call up each trace if required. The end result is a fully automated process from assay interpretation to reporting, with full and continuous QA data logging.
The company has successfully collaborated with a number of laboratories both here and abroad, including the Department of Laboratory Medicine at the University of Washington, where over 4,000 cytomegalovirus samples were analysed. Diagnostics.ai software was found to be highly accurate and efficient at quantifying CMV viral loads and that the automatic data analysis reduced errors and required less hands-on analysis time.
Another study was carried out with the West of Scotland Specialist Virology Centre at the Gartnavel General Hospital in Glasgow. This showed that results from different assays analysed and run on different types of thermal cyclers were in complete agreement with manual methods. However, interpretation was significantly faster using the new software, with the additional benefits of reducing turn-around times and staff costs.
A summary of some of the benefits of diagnostics.ai technology are shown in table 1.
For further information, please contact:
Dr Mel Smith:
melvyn [dot] smith [at] nhs [dot] net

